All published articles of this journal are available on ScienceDirect.
Vitamin D Receptor Gene Polymorphisms and the Risk of Chronic Hepatitis C Related Hepatocellular Carcinoma in Egyptian Population
Abstract
Background:
Small percentage of hepatitis C (HCV) patients develop hepatocellular carcinoma (HCC) during their lifetime, suggesting that genetic factors might modulate HCC development. Numerous variations on the vitamin D receptor gene (VDR) have been recognized in human cancers. The majority of them cause VDR to be unable to bind to 1, 25-OH-D. The aim of the present work was to investigate the relation of VDR FokI (rs2228570), BsmI (rs3782905) and ApaI (rs7975232) gene polymorphisms and the risk of HCC development in chronic HCV Egyptian patients.
Methods:
A total of 311 Egyptian patients were enrolled for this study. They were divided into 3 groups: 103 patients with liver Cirrhosis, 107 patients with HCC and 101 normal healthy subjects as the control group. Human genomic DNA Extraction was carried out using QIAamp® DNA Blood Mini Kit (QIAGEN) Genotyping of VDR ApaI (rs7975232) single nucleotide polymorphism (SNP) was carried out using real-time PCR TaqMan allelic discrimination assay with allele-specific designed fluorescent MGB probes.
Results:
Patients with HCC had a higher frequency of ApaI CC genotype (P=0.035) CI (0.031-0.038). Patients with HCC carried a higher ratio of ApaI CC genotype compared to those with liver cirrhosis (x2=5.4 and P = 0.03) or controls (x2=6.8 and P = 0.01). Univariate analysis revealed that age, lower platelet count (<150×103/μL), higher AFP (>100 ng/ml), and ApaI CC genotype were the factors significantly associated with the development of HCC. Stepwise logistic regression analysis showed that all were independent predictors.
Conclusion:
ApaI CC VDR gene mutation is an independent risk factor for HCC development in Egyptian Cirrhotic HCV patients.
1. INTRODUCTION
Hepatocellular carcinoma (HCC) is a major worldwide malignancy. Hepatocarcinogenesis is a complex process of variable etiologies mainly secondary to viral hepatitis infection with hepatitis B virus (HBV) and hepatitis C virus (HCV) [1].
Egypt suffers the highest prevalence of HCV infection, overwhelming their local health services. Chronic HCV infection is characterized by a high rate of progression to fibrosis, chronic hepatitis, and eventually cirrhosis and ultimately to HCC [2]. Some of the host- and viral-related pathogenetic mechanisms of hepatocarcinogenesis are still not well known. There is a continued need for the identification of hepatocarcinogenesis genetic risk factors for early identification and preventative interventions [3].
Vitamin D is a systemic hormone that plays an important role in the metabolism of the skeleton and helps mainly the regulation of host immune responses and fibrogenesis. It has been established that mutations in genes involved in response to hormones, their metabolism or actions may affect the outcome of the disease and thus act as modifiers [4]. This could explain its role in carcinogenesis through vitamin D receptor gene (VDR) polymorphism that made it unable to bind to 1, 25-OH-D [5, 6].
Vitamin D insufficiency has been identified in different populations worldwide as well as its correlation with the presence of various malignancies, including HCC [6]. The VDR gene is a highly polymorphic gene located on chromosome 12q12–q14 [7]. Various single-nucleotide polymorphisms (SNPs) were identified. These genetic variations in the VDR gene could explain the inter-individual variations in rates of vitamin D synthesis and activation in the skin, liver and kidney, transport, metabolism and degradation. These phenotypic variations could influence the anti-tumor effect of vitamin D through the change in cell behaviors such as proliferation, differentiation and apoptosis. Moreover, cumulative data shows the relation between the VDR gene variants and increased risk of cancers including breast [8], prostate [9], and colorectal [10]. VDR polymorphisms were investigated in various chronic liver diseases, including primary biliary cirrhosis and autoimmune hepatitis [11]. So, the aim of this work was to investigate the relation of VDR FokI (rs2228570), BsmI (rs3782905) and ApaI (rs7975232) gene polymorphism and the risk of HCC development in chronic HCV Egyptian patients.
2. METHODS
This cross-sectional multi-center study was carried out between May 2018 and June 2019. A total of 468 were recruited for participation from University Hospitals in Kafrelsheikh, Banha, Minia, Cairo and Alexandria Medical research Institute Tanta outpatient hepatology clinics and inwards, 258 were excluded and 210 Egyptian participants were enrolled. They were divided into two groups; chronic HCV patients group I included 107 patients with cirrhosis and 103 patients with HCC group II. A third control group was included, 101 normal healthy subjects with matched age and sex from blood bank donors were enrolled. The study was approved by the Ethics Committee of each Faculty of Medicine in accordance with the Helsinki Declaration of 1975. In addition, written informed consent was obtained from each participant prior to their inclusion in the study.
2.1. Inclusion Criteria
All patients groups were previously confirmed for seropositive HCV antibody and HCV RNA by MX 3000. Liver cirrhosis was diagnosed by abdominal ultrasound. Diagnosis of HCC was based on at least one positive HCC image on Computed Tomography (CT) or Magnetic Resonance Imaging (MRI), with or without elevated serum AFP level.
2.2. Exclusion Criteria
Positive hepatitis B surface antigen patients, positive HIV patients or patients with any other causes of cirrhosis (e.g., Alcoholism, severe nonalcoholic liver disease with metabolic syndrome, autoimmune hepatitis or primary biliary cirrhosis) were excluded from this study. Those with a prior diagnosis of HCC who underwent any therapeutic interventions before were also excluded.
All subjects were subjected to thorough history-taking including special habits like smoking and clinical examination.
2.3. Sample Collection and Laboratory Analysis
10 ml of venous blood was collected after overnight fasting and divided into 4 sample tubes. First 4 ml in a serum separator tube for blood chemistry, including liver function tests; aspartate aminotransferase (AST), alanine aminotransferase (ALT), total bilirubin and albumin, kidney function tests; creatinine and urea by using a Beckman CX4 chemistry analyzer (NY, USA, supplied by the Eastern Co. For Eng.& Trade-Giza, Egypt) [12]. Serum AFP was assayed using Abbott, Axyam (USA, Supplied by Al-Kamal company Cairo, Egypt). The second 2 ml was separated on EDTA vacutainer tubes for complete blood picture by phoenix 3300. The third 2 ml was separated in sodium for prothrombin time PT/ international normalized ratio INR measurements by KCL Delta. The last 2 ml was withdrawn in separate EDTA vacutainer tubes for VDR polymorphism detection. They were stored at –80ºC until the DNA extraction procedure.
2.4. Genetic Analysis of SNPs
2.4.1. Human Genomic DNA Extraction
The extraction was carried out using QIAamp® DNA Blood Mini Kit (QIAGEN) according to the manufacturer's instructions [13]. The concentration of the extracted DNA was measured using the Nano Drop® (ND-1000) Spectrophotometer (Nano Drop Technologies Inc., Washington, USA). The ratio of absorbance of extracted DNA at 260 /280 nm was 1.7–1.9.
2.4.2. Genotyping
Genotyping of VDR FokI (rs2228570), BsmI (rs3782905) and ApaI (rs7975232) single nucleotide polymorphisms (SNP) was carried out independently using real-time polymerase chain reaction PCR TaqMan allelic discrimination assay with allele-specific designed fluorescent MGB probes, obtained from Applied Biosystems (Applied Biosystems-Life Technologies, Carlsbad, California, USA) [14].
2.4.3. Statistical Analysis
All analyses were carried out using IBM SPSS Statistics for Windows, version 20.0 (IBM Corp., Armonk, N.Y., USA). Continuous data are expressed as mean ± standard deviation, and the categorical data are expressed as number (percentage). Comparisons of differences in the categorical data between groups were performed using the chi-square test. Distributions of continuous variables were analyzed by the Student’s t-test or one-way ANOVA test with the least significant difference (LSD) post-hoc correction between groups where appropriate. Stepwise logistic regression analysis was performed to assess the influence of each factor on the risk of developing HCC. All tests were 2-tailed, and a p-value of less than 0.05 was considered statistically significant.
3. RESULTS
Total 311 participants of Egyptian origin were enrolled in this study. They were divided into 3 groups: 103 patients with HCV liver Cirrhosis, 107 patients with HCV related HCC and 101 normal healthy subjects as a control group.
3.1. Demographical and Laboratory Features of the Studied Population
Table 1 shows the baseline demographic and laboratory features of the studied groups. The mean age of HCC patients was significantly higher than those with liver cirrhosis and controls (P < 0.001). As regards sex and BMI, no statistically significant differences were observed between these groups. The smoking percentage of HCC patients was significantly higher than the other two groups (P =0.023). In addition, patients with HCC had high statistically significant different laboratory results for AST, ALT, T-Bil, ALB, creatinine, platelet count, INR and AFP (P<0.001).
3.2. Genotypic and Allelic Distribution of VDR SNPs (FokI, Bsm1and ApaI) among the Three Studied Groups.
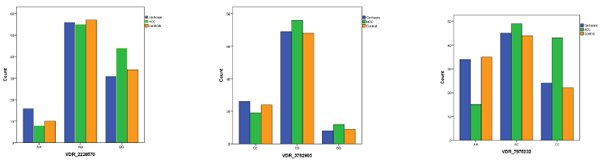
Patients carrying the corresponding ApaI CC genotype had a higher prevalence (48.3%) of HCC than those with CA (35.5%) or AA type (19.7%) (x2=18.5 and P <0.001). Patients with HCC carried a higher ratio of ApaI CC genotype compared to those with liver cirrhosis (x2=5.4 and P = 0.03) or controls (x2=6.8 and P = 0.01). CI (0.031-0.038). For the FokI and BsmI polymorphisms, no significant associations were found (Table 2) (Fig. 1).
| Parameter |
Liver Cirrhosis (n = 103) Mean ± SD |
HCC (n = 107) Mean ± SD |
Control (n = 101) Mean ± SD |
P-value |
|---|---|---|---|---|
| Age(years) | 54.5±9.74 | 59.19±7.69 | 55.96±8.68 | < 0.001* |
| Male gender, n (%) | 59 (57.3) | 60 (56.1) | 59 (58.4) | 0.477 |
| Smoking, n (%) | 63(61.1) | 72(67.3) | 61(60.3) | 0.023* |
| BMI (kg/m2) | 26.63±2.9 | 26.36±2.1 | 25.57±3.9 | 0.07 |
| AST (U/L) | 60.6±30.2 | 102.1±53.9 | 32.5±6.6 | < 0.001* |
| ALT (U/L) | 44.2±25.4 | 59.3±28.1 | 30±5.5 | < 0.001* |
| T-Bil (mg/dL) | 3.9±1.3 | 6.27±2.5 | 0.8±0.3 | < 0.001* |
| ALB(g/L) | 2.5±0.8 | 2.6±0.5 | 3.9±0.2 | < 0.001* |
| Creatinine(mg/dL) | 1.54±1.14 | 2.26±1.79 | 0.96±0.16 | < 0.001* |
| Platelet (103/μL) | 140.5±72 | 114±41.6 | 286.2±90.1 | < 0.001* |
| INR | 1.4±0.3 | 1.6±0.7 | 1±0.1 | < 0.001* |
| AFP (ng/ml) | 135.2±20 | 1567±202.5 | 5.72±1.7 | < 0.001* |
|
Gene SNP/rs No. |
Allele | Genotype Distribution | P-value | Allelic OR* (95% CI) | ||
|---|---|---|---|---|---|---|
| Liver Cirrhosis N=103 (%) |
HCC N=107(%) |
Control N=101(%) |
- | - | ||
| - | GG | 34 (33.7) | 44 (41.1) | 31 (30.1) | - | - |
| FokI (rs2228570) |
GA | 57 (56.4) | 55 (51.4) | 56 (54.4) | 0.22 | 1.45 (0.8-2.6) |
| - | AA | 10 (9.9) | 8 (7.5) | 16 (15.5) | 0.03 | 2.8 (1.08-7.45) |
| MAF | G vs. A allele | 125: 77 | 143: 71 | 118: 88 | - | - |
| P value of HWE | - | - | - | - | 0.26 | - |
| CC | 24 (23.8) | 19 (17.8) | 26 (25.2) | - | - | |
| Bsm1 (rs3782905) |
CG | 68 (67.3) | 76 (71) | 69 (67.0) | 0.3 | 1.5 (0.77-2.96) |
| GG | 9 (8.9) | 12 (11.2) | 8 (7.8) | 0.3 | 2 (0.7-5.9) | |
| MAF | G vs. C allele | 86:116 | 100:114 | 89: 121 | - | - |
| P value of HWE | - | - | - | - | 0.39 | - |
| AA | 35 (34.7) | 15 (14) | 34 (33) | - | - | |
| ApaI (rs7975232) |
AC | 44 (43.6) | 49 (45.8) | 45 (43.7) | 0.2 | 2.5(1.19-5.12) |
| CC | 22 (21.8) | 43 (40.2) | 24 (23.3) | <0.001 | 4.1(1.86-8.92) | |
| MAF | C vs. A allele | 88:114 | 135:79 | 93:113 | - | - |
| P value of HWE | - | - | - | - | 0.8 | - |
3.3. Factors Associated with Developing HCC by Logistic Regression Analysis
Univariate analysis revealed that age, lower platelet count (<150× 103/μL), higher AFP (>100 ng/ml) and ApaI CC genotype were factors significantly associated with developing HCC. Stepwise logistic regression analysis showed that all were independent predictors (Table 3).
| Univariate Analyses | Stepwise Multivariate Analyses | |||
|---|---|---|---|---|
| - | Odds Ratio (95% CI) |
P-value | Odds Ratio (95% CI) |
P-value |
| Age (per 1-year increase) | 1.05(1.03-1.11 | <0.001 | 1.1(1.1-1.15) | <0.001 |
| Male gender | 0.9(0.6-1.6) | 0.9 | - | - |
| BMI | 0.9(0.9-1.1) | 0.4 | - | - |
| Smoking | 1.3(0.7-2.3) | 0.3 | - | - |
| Platelet (<150 × 103/μL) | 2.4 (1.27-4.35) | 0.006 | 4.8(2.2-10.3) | <0.001 |
| AFP (>100 ng/ml) | 2.4(1.4-4.3) | 0.002 | 3(1.5-6) | 0.001 |
| FokI GG type | 2.1(1.08-7.5) | 0.06 | - | - |
| BsmI GG type | 2(0.7-6) | 0.2 | - | - |
| ApaI CC type | 4.1(1.8-8.9) | <0.001 | 5.6(2.3-13.9) | <0.001 |
4. DISCUSSION
HCV-related HCC is a major health problem, with significant mortality and morbidity rates being the fifth most common cancer and the second leading cause of cancer-related death worldwide [15-35]. Nevertheless, only a fraction of infected patients develop HCC during their lifetime, suggesting that genetic factors might modulate HCC development [16]. Consequently, the identification of additional genetic factors affecting transcription of specific regulatory genes could help to select high-risk populations. So, the aim of the present work was to investigate the possible association between the VDR gene polymorphisms (FOKI rs 2228570, BsmI rs3782905 and ApaI rs7975232) and HCC in the Egyptian population with chronic HCV infection.
The present work demonstrated that smoking among HCC patients was significantly higher than those with liver cirrhosis and controls. In agreement with our result, a recent meta-analysis has shown a significant increase in relative risk of HCC in smokers with HCV compared with nonsmokers with HCV, with relative risks of 23 and 7.9 respectively [17] this probably could be explained by the fact that HCC risk is linked to lifestyle factors, such as smoking which accelerate progression to HCC in HCV patients likely via increased oxidative stress.
In the present study, we found a significantly higher level of serum transaminases (ALT and AST) in the patients’ groups than in control. AST and ALT are excellent markers of hepatocellular injury, especially in the setting of chronic HCV infection [18]. Also, we demonstrated impairment of both secretory (as evident by increased bilirubin level) and synthetic (as evident by decreased albumin level and prolongation of prothrombin time) functions of the liver, which were significantly obvious in the patients’ groups than in the control group. Also, the platelets count was significantly lower in the patients’ groups than in the control group. These findings were due to the necro-inflammatory effect of chronic HCV infection. Furthermore, due to the tumor mass cholestatic and necrotic effects of HCC on the liver tissue, AST and serum bilirubin level were significantly higher in the HCC group than liver cirrhosis group [19].
AFP can be mildly elevated in those with regenerating nodules in viral cirrhosis, giving more false-positive results. However, a rising AFP over time is more accurate in the diagnosis of HCC, even if the level does not reach 400 ng/ml [19]. In agreement with the previously mentioned data, the present work demonstrated a significantly higher level of AFP in the patients’ group than the control group. Furthermore, the level of AFP was significantly higher in HCC group than in liver cirrhosis group. Also, in 2018 the American association of study of liver diseases guidelines stated that surveillance should be accomplished by ultrasound with or without AFP, as it is “not possible to determine which type of surveillance test, US [ultrasound] alone or the combination of US plus AFP, leads to greater improvement in survival” [20].
In the present work, we demonstrated the prevalence of VDR gene polymorphism (FokI rs 2228570, BsmI rs3782905 & ApaI rs7975232) in HCC and liver cirrhosis, and we found that patients with HCC had a higher frequency of ApaI CC genotype (40.2%) than in those with liver cirrhosis (23.35%) and the control group (21.8%). Moreover, logistic regression analysis showed that ApaI CC genotype was an independent risk factor for HCC making ApaI CC genotype a potential risk factor for HCC in patients with HCV liver cirrhosis. On the other hand, the other ApaI variant AA and AC genotypes were found to be associated with a significantly increased risk of renal cell carcinoma compared with the CC genotype [21].
In agreement with our results, Hung et al. [22] suggested a significant association of VDR ApaI rs 7975232 polymorphism with the development of HCC in chronic HCV infection. In his hospital-based study, patients with HCC also had a higher frequency of ApaI CC genotype. Other two studies have reported the significant relationship between VDR gene polymorphisms and HCC development in patients with chronic HCV infection [23, 24].
In another Egyptian study [25] on chronic HCV patients, in accordance with our result, Rawhia et al. found that VDR Apa-I gene polymorphism could be associated with an increased risk of HCC development in chronic hepatitis C virus-infected patients. Also, Galal et al. [26] had found that the carriage of the ApaI CC genotype was an independent predictor for HCC in HCV-related liver cirrhosis.
Although, Falleti et al. [27], in their study of VDR genetic polymorphisms in transplantation patients of variable etiologies, found that A strong association was observed between carriage of the BAT A-T-C and G-T-T haplotypes and HCC only in alcoholic liver disease rather than to those with viral cirrhosis. Triantos et al. [28] showed that the carriage of VDR ApaI AA genotype was associated with more severe liver disease compared to ApaI CC/CA genotypes. This contradiction could be justified by the difference in the inclusion criteria as the authors investigated patients with chronic liver disease of variable causes (viral, autoimmune, cryptogenic, etc.) and not complicated by HCC on top, while in our cohort, we selected only the HCV-related cirrhotic patients.
Moreover, in an Italian study [29] on chronic HCV patients treated with direct acting antivirals (DAAs), the authors elucidated vitamin D pathway gene SNPs and HCC relationship in the Italian population. They showed that HCC risk factors were age, ribavirin administration, IL28Brs12979860CC and previous treatments; VDR FokI CC, sex and insulin resistance were protective factors. Although the studied SNPs were different from our work.
Our study has the advantage of being multicentric, covering most of the geographical areas of Egypt, but the study has its own limitations. The relatively small number of patients included but this can be explained by the pilot nature of the study and trial to cover more geographical areas. Our study is still a cross-sectional study, and we are now following our cirrhotic patients that have VDR ApaI CC polymorphism to confirm the causal relationship.
CONCLUSION
ApaI CC VDR gene mutation is an independent risk factor for HCC development in Egyptian Cirrhotic HCV patients.
LIST OF ABBREVIATIONS
| ALT | = Alanine aminotransferase |
| AST | = Aspartate aminotransferase |
| CT | = Computed Tomography |
| DAAs | = Direct Acting Antivirals |
| HBV | = Hepatitis B Virus |
| HCC | = Hepatocellular Carcinoma |
| HCV | = Hepatitis C Virus |
| MRI | = Magnetic Resonance Imaging |
| SNPs | = Single-Nucleotide Polymorphisms |
| VDR | = Vitamin D Receptor |
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
The study was approved by the Ethics Committee of Kafrelsheikh, Tanta, Banha, Minia, Cairo Faculties of Medicine and the Alexandria Medical Research Institute.
HUMAN AND ANIMAL RIGHTS
No animals were used in this research. All human research procedures followed were in accordance with the ethical standards of the committee responsible for human experimentation (institutional and national), and with the Helsinki Declaration of 1975, as revised in 2013.
CONSENT FOR PUBLICATION
Written informed consent was obtained from each participant prior to their inclusion in the study.
AVAILABILITY OF DATA AND MATERIALS
The data that support the findings of this study are available from (Biochemistry Department, National Hepatology & Tropical Medicine Research Institute, Cairo University) but restrictions apply to the availability of these data, which were used under license for the current study, and so are not publicly available. Data are, however, available from the authors upon reasonable request and with permission of the Biochemistry Department, National Hepatology & Tropical Medicine Research Institute.
FUNDING
None.
CONFLICT OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
The authors would thank all the nursing staff and laboratory technicians who helped in this work.


