All published articles of this journal are available on ScienceDirect.
Association Between Vitamin D Receptor Gene Polymorphisms and Helicobacter Pylori Infection
Abstract
Background & Aims:
Human genetic polymorphism has been reported in the susceptibility and clinical development of infection. In this regard, this study aimed to investigate the link between Vitamin D Receptor (VDR) gene polymorphism and H. pylori infection.
Materials and Methods:
This cross-sectional study was conducted on 224 adult patients with upper gastrointestinal symptoms who underwent an upper gastrointestinal endoscopy between July 2017 and May 2019 in two major university hospitals. All patients were evaluated for helicobacter pylori infection. Two gastric antral biopsy specimens were taken from each patient. One of those Biopsy specimens was evaluated for histopathology examination and the other one was immersed in a saline solution ready for genomic DNA extraction.
Results:
There were statistically significant differences between different genotypes of VDR rs7975232 polymorphism between H. pylori infected and non-infected groups (CC was higher in H. pylori negative and AC and AA were the most common in H. pylori positive group). There were statistical differences between different genotypes of VDR rs2228570 between H. pylori infected and non-infected groups (TT was higher in H. pylori negative and CT and CC were the most common in H. pylori positive group). Regarding VDR rs 7975232 gene polymorphisms; the (A) allele was significantly higher H. pylori infected, while (C) allele was significantly higher in uninfected patients. Regarding VDR rs 2228570 gene polymorphisms; the (C) allele was significantly higher H. pylori infected, while (T) allele was significantly higher in uninfected patients.
Conclusion:
There is a possible association between the FokI and Apal VDR polymorphism and H. pylori infection.
1. INTRODUCTION
Warren and Marshall discovered Helicobacter pylori (H. pylori) in 1983 and reported it in 1984 and in 2005, and were awarded the Noble prize [1].
H. pylori are prevalent throughout the world but are more prevalent in developing countries [2, 3]. This infection is also present more in elderly persons than adolescents [4].
H. pylori infection causes many gastric diseases, such as chronic gastritis, peptic ulcer disease, gastric cancer and gastric Mucosa-Associated Lymphoid Tissue (MALT) lymphoma [5].
Helicobacter pylori are spiral-shaped Gram-negative bacteria. Due to the properties of the gastric mucosa, H. pylori adhesion and survival in such an environment depend on many mechanisms and specific virulence factors [6]. Strains that live in the gastric mucosa and how they interact with the host’s immune system determine the H. pylori long-term colonization with chronic gastritis induction [7].
Also, human genetic polymorphisms contribution to the susceptibility and clinical evolution of the infection was reported. They can produce different immune system mediators' expression levels which are in relation to infection rates, H. pylori eradication, as well as colonization by more virulent strains, gastric inflammation enhancement and cell damage [8]. An improvement in Vitamin D Receptor (VDR) expression in gastric epithelia was proposed to be induced by H. pylori infection. In vitro, the active form of vitamin D, 1a-25-dihydroxyvitamin D3, was obtained by stimulating the cathelicidin antimicrobial peptides and decreasing inflammatory cytokine and chemokine levels, forming immune modulators properties against this pathogen [9].
The location of the gene responsible for VDR expression is on the 12q13.11 chromosome and comprises a region of >60 kb of DNA [10]. The presence of Single Nucleotide Polymorphisms (SNP) was reported on this gene. The Restriction Fragment Length Polymorphism (RFLP) for FokI, BsmI, ApaI and TaqI restriction enzymes is used nowadays in genetic susceptibility studies of infectious diseases [11]. TaqI polymorphism (rs731236; T>C) results from the change of thymine to cytosine (ATT to ATC). The FokI polymorphism (rs2228570; C>T), characterized by the change of thymine to cytosine (ATG to ACG) [12].The noncoding regions VDR gene polymorphisms are the BsmI polymorphism (rs1544410; G>A) where guanine is changed to adenine (GCG to GCA) and ApaI polymorphism (rs7975232, A>C) where cytosine is changed to adenine [13]. Low vitamin D levels were associated with lower H. pylori eradication rate. Therefore, the vitamin D signal immune function impairment, which may lead to inadequate immune response, is a potential pathogenic mechanism that explains the observed association between vitamin D status and eradication rates [14].
Vitamin D plays a pivotal role in both bones and non-bone organs [15]. Both vitamin D paracrine and autocrine biological activities are exerted by binding to its intracellular receptor known as Vitamin D Receptor (VDR). The widespread VDR expression in numerous tissues, including the kidney, heart, muscle, breast, colon, prostate, brain and immune cells, makes it as a natural target of modulation in disease pathogenesis including dermal disorders, metabolic syndrome, variety of cancers and renal transplant [15, 16].
There is an association between vitamin D status and H. pylori eradication rates, as lower 25-OHD was associated with H. pylori eradication failure [17].
In this regard, this study aimed to investigate the link between VDR gene polymorphism in H. pylori infection.
2. MATERIALS AND METHODS
The study protocol was performed according to the ethical guidelines of the Helsinki Declaration and was approved by the ethical committee of Tanta University Faculty of Medicine. Written informed consent was signed by all patients participating in the study.
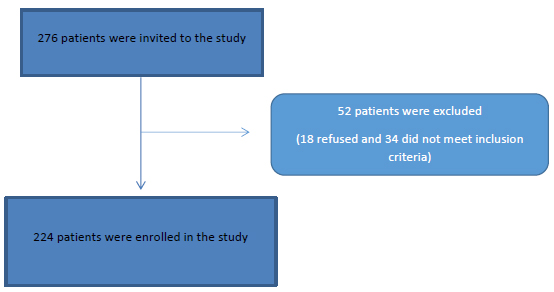
This cross-sectional study was conducted on 276 adult patients attending the internal medicine department, Cairo University Hospitals and Tropical medicine department, Tanta University with upper gastrointestinal symptoms who underwent an upper gastrointestinal endoscopy between July 2017 and May 2019 Fig. (1).
After previous evaluations due to the presence of symptoms in the upper gastrointestinal tract, patients who showed dyspeptic disorders and who were subsequently diagnosed with chronic gastritis were considered for the study.
Subjects who had received anti-Helicobacter therapy within the last 6 months, or used a proton pump inhibitor within the last 30 days, pregnant females, patients with a history of alcohol abuse or with HIV infection were excluded from the study.
All patients underwent the following:
A). Anthropometric Measurements
Body Mass Index (BMI) was calculated as body weight in kilograms divided by body height squared in meters
B). Laboratory Analysis
CBC, AST, ALT, T. Bilirubin, D. Bilirubin, S. albumin, Alkaline phosphatse, fasting blood sugar, AFP, S. creatinine and H. pylori antigen in stool.
Helicobacter pylori Infection Test: The diagnosis of H. pylori infection was done by ELISA technique using the IBL International Kit (Flughafenstrasse, Hamburg, Germany), and Tecan Spectra ELISA reader (supplied by Tecan Group Ltd. Switzerland); With 3 ng/ml as a cut-off value.
C). Upper Endoscopy
Two gastric antral biopsy specimens were taken from each patient. One of those Biopsy specimens for histopathology examination and the other one was immersed in a saline solution (0.9%) ready for genomic DNA extraction.
DNA extraction and genotyping: The DNA extraction was carried out using QIAamp® DNA Blood Mini Kit (QIAGEN GmbH, Hilden, Germany) (Schur, Bjerke et al. 2001) according to the manufacturer's instructions. The concentration of the extracted DNA was measured using the Nano Drop® (ND-1000) Spectrophotometer (Nano Drop Technologies Inc., Washington, USA). The ratio of absorbance of extracted DNA at 260 /280 nm was 1.7-1.9.
Genotyping vit D receptor SNPs: Genotyping of VDR FokI (rs2228570) was carried out, where the substitution of thymine to cytosine (ATG to ACG) changed the protein structure of the VDR receptor. The presence of the mutant allele and ApaI (rs7975232) where (A>C) indicates that cytosine was changed to adenine and Single Nucleotide Polymorphisms (SNP) were carried out independently using real-time PCR TaqMan allelic discrimination assay with allele-specific designed fluorescent MGB probes obtained from Applied Biosystems (Applied Biosystems-Life Technologies, Carlsbad, California, USA) [18].
Statistical analysis of the data: Data were fed to the computer and analyzed using IBM SPSS software package version 20.0. (Armonk, NY: IBM Corp). Qualitative data were described using number and percent. Quantitative data were described using range (minimum and maximum), mean, standard deviation and median. The significance of the obtained results was judged at the 5% level. The Chi-square test or Monte Carlo which was used for categorical variables, while student t-test was used for numerical variables to compare between different groups. Regression analysis was used to test the effect of different factors on VDR rs 7975232 and VDR rs 2228570.
3. RESULTS
The study was conducted on 224 patients with the mean age (39.76 ± 8.440). Females were 141(63%) and males were 83(37%) among patients and there was no significant difference between the different genotypes regarding sex. Baseline data of the included patients are shown in Table 1 and Table 3.
Regarding AST, ALT, s. albumin and Hemoglobin level, they were significantly higher in h.pylori infected patients than the non-infected group. However, blood glucose, serum creatinine and BMI were significantly higher in the non-infected group than H. pylori infected patients Table 2.
Regarding VDR rs 7975232 gene polymorphisms, CC, AC and AA genotypes distribution among the total studied subjects were 42%, 33% and 25%, respectively. Regarding VDR rs 7975232 gene polymorphisms; CC, AC and AA genotypes distribution were 16.5% &25.4%, 12.9% &20.1% and 7.6% &17.4%, among males and females, respectively).
Regarding VDR rs 2228570 gene polymorphisms, TT, CT and CC genotypes distribution among the total studied subjects were 32.1%, 42% and 25.9%, respectively. Regarding VDR rs 2228570 gene polymorphisms, TT, CT and CC genotypes distribution were 12.9% &19.2%, 16.5% &25.4% and 7.6% &18.3%, among males and females, respectively) as shown in Table 3.
H. pylori infection was positive in 127 (56.7%) patients. There were statistical differences between different genotypes of VDR rs 7975232 between H. pylori infected and non-infected groups (CC was higher in H. pylori negative (64.9%) and AC and AA were the most common in H. pylori positive group (42.5% and 33.1%, respectively). This detail is given in Table 4.
There were statistical differences between different genotypes of VDR rs2228570 between H. pylori infected and non-infected groups (TT was higher in H. pylori negative (46.6%) and CT and CC were the most common in H. pylori positive group (40.9% and 37.8%, respectively).
Regarding VDR rs 7975232 gene polymorphisms, alleles distribution among the total studied subjects were (C and A were 75% and 58%, respectively), while (A) allele was significantly higher H. pylori infected (75.6&), while (C) allele was significantly higher in uninfected patients (85.6%). Regarding VDR rs 2228570 gene polymorphisms, alleles distributions among the total studied subjects were (T and C) 74.1% and 67.9%, respectively, while (C) allele was significantly higher H. pylori infected (78.7%), while (T) allele was significantly higher in uninfected patients (89.7%). This is demonstrated in Table 5.
After regression analysis; blood glucose was the only independent risk factor of rs 7975232 Table 6. Blood glucose and s. albumin were the only negatively independent risk factors of rs 2228570 Table 7.
| Variables | N (224) | Minimum | Maximum | Mean ± SD |
|---|---|---|---|---|
| Age (years) | 224 | 20 | 65 | 39.8±8.4 |
| AST (IU/L) | 224 | 10 | 41 | 28.1±7.3 |
| ALT(IU/L) | 224 | 10 | 58 | 30.9±9.0 |
| T. Bilirubin (mg/dL) | 224 | 0.5 | 2.8 | 0.9±0.4 |
| D. Bilirubin(mg/dL) | 224 | 0.1 | 0.9 | 0.2±0.1 |
| Albumin (g/dL) | 224 | 2.7 | 5.0 | 3.8±0.4 |
| Glucose (mg/dL) | 224 | 66 | 398 | 165.8±90.4 |
| A FP (ng/ml) | 224 | 3.0 | 91.0 | 10.3±7.6 |
| Hb (g/dL) | 224 | 8.0 | 15.0 | 11.4±1.9 |
| S.Creatinine(mg/dL) | 224 | 0.6 | 1.6 | 1.0±0.2 |
| BMI (Kg/m2) | 224 | 15 | 45 | 30.6±7.3 |
| Platlets | 224 | 165 | 450 | 304.1±80.1 |
| Alkal.Phosph. (IU/L) | 224 | 5 | 223 | 101.3±34.5 |
| - | H. pylori | P-value | |
|---|---|---|---|
| Positive (Mean+SD) | Negative (Mean+SD) |
||
| Age | 39.47+8.77 | 40.14+8.02 | 0.556 |
| AST | 30.57+6.30 | 24.87+7.25 | 0.000* |
| T. Bilirubin | 0.93+0.25 | 0.97+0.54 | 0.415 |
| D. Bilirubin | 0.22+0.08 | 0.22+0.19 | 0.675 |
| ALB | 3.89+0.38 | 3.70+0.45 | 0.001* |
| Glucose | 92.81+12.88 | 261.28+50.01 | 0.000* |
| AFP | 10.78+9.56 | 9.60+3.57 | 0.250 |
| Hb | 12.30+1.77 | 10.23+1.26 | 0.000* |
| Creatinine | 0.95+0.15 | 1.09+0.27 | 0.001* |
| BMI | 28.71+7.10 | 33.03+6.82 | 0.002* |
| Plt | 306.26+79.78 | 301.33+80.77 | 0.649 |
| ALP | 101.01+34.77 | 101.56+34.38 | 0.906 |
| VDR Gene Polymorphism |
Genotypes | Gender | Total (224) | Significance | |
|---|---|---|---|---|---|
| Male (83=37%) | Female (141=63%) | ||||
| rs 7975232 | CC | 37 (16.5%) | 57 (25.4%) | 94 (42%) | 0.176 |
| AC | 29 (12.9%) | 45 (20.1%) | 74 (33%) | ||
| AA | 17 (7.6%) | 39 (17.4%) | 56 (25%) | ||
| rs 2228570 | TT | 29 (12.9%) | 43 (19.2%) | 72 (32.1%) | 0.128 |
| CT | 37 (16.5%) | 57 (25.4%) | 94 (42%) | ||
| CC | 17 (7.6%) | 41 (18.3%) | 58 (25.9%) | ||
4. DISCUSSION
This study aimed to investigate the link between VDR gene polymorphism in H. pylori-infected patients with gastric symptomatology.
The study was conducted on 224 patients with the mean age (39.76 ± 8.440). Females were 141(63%) and males were 83 (37%) among patients and there was no significant difference between the different genotypes regarding sex.
H. pylori infection was present in 127 (56.7%) patients (Table 3). H. pylori infection showed 58% prevalence in patients with chronic gastritis [19], which is nearly similar to our results.
There were statistical differences between different genotypes of VDR rs 7975232 (Apal) between H. pylori infected and non-infected groups. CC was higher in H. pylori negative (64.9%) and AC and AA were the most common in H. pylori positive group (42.5% and 33.1%, respectively). There was statistical difference between different genotypes of VDR rs2228570 (FokI) between H. pylori infected and non-infected groups.TT was higher in H. pylori negative (46.6%) and CT and CC were the most common in H. pylori positive group (40.9% and 37.8%, respectively).
| VDR | H. pylori (224) | p- value | |||
|---|---|---|---|---|---|
| Positive 127(56.7%) |
Negative 97(43.3%) |
Total | |||
| rs 7975232 | CC | 31(24.4%) | 63(64.9%) | 94(42%) | 0.000* |
| AC | 54(42.5%) | 20(20.6%) | 74(33%) | ||
| AA | 42(33.1%) | 14(14.4%) | 56(25%) | ||
| rs 2228570 | TT | 27(21.3%) | 45(46.4%) | 72(32.1%) | 0.001* |
| CT | 52(40.9%) | 42(43.3%) | 94(42%) | ||
| CC | 48(37.8%) | 10(10.3%) | 58(25.9%) | ||
| Genotype | Alleles | H. pylori (224) | p- value | ||
|---|---|---|---|---|---|
| Positive 127(56.7%) |
Negative 97(43.3%) |
Total | |||
| rs 7975232 | C | 85 (66.9%) | 83 (85.6%) | 168 (75%) | 0.002 |
| A | 96 (75.6%) | 34 (35.1%) | 130 (58%) | ||
| rs 2228570 | T | 79 (62.2%) | 87 (89.7%) | 166 (74.1%) | 0.000 |
| C | 100 (78.7%) | 52 (53.6%) | 152 (67.9%) | ||
| - | Unstandardized Coefficients | Standardized Coefficients | - | - | |
|---|---|---|---|---|---|
| Model | B | Std. Error | Beta | T | Sig. |
| (Constant) | 1.940 | .344 | 5.635 | .000 | |
| Age | -.002 | .006 | -.017 | -.265 | .792 |
| Ast | -.253 | .393 | -.055 | -.644 | .520 |
| Alt | .448 | .259 | .148 | 1.726 | .086 |
| T BIL | -.039 | .182 | -.019 | -.216 | .829 |
| D BIL | .006 | .171 | .003 | .036 | .971 |
| Alb | -.146 | .151 | -.072 | -.969 | .334 |
| Glu | -.544 | .152 | -.338 | -3.585 | .000 |
| a FP | .068 | .116 | .040 | .585 | .559 |
| Hb | .025 | .162 | .014 | .152 | .879 |
| Creatine | .046 | .135 | .024 | .338 | .735 |
| Bmi | .027 | .124 | .015 | .216 | .829 |
| Alp | -.254 | .176 | -.098 | -1.440 | .151 |
| Gender | .102 | .110 | .061 | .923 | .357 |
| - | Unstandardized Coefficients | Standardized Coefficients | - | - | |
|---|---|---|---|---|---|
| Model | B | Std. Error | Beta | T | Sig. |
| (Constant) | 2.302 | .321 | 7.179 | .000 | |
| Age | -.005 | .006 | -.054 | -.840 | .402 |
| AST | -.491 | .366 | -.113 | -1.341 | .181 |
| ALT | .328 | .241 | .114 | 1.357 | .176 |
| T.Bilirubin | .268 | .169 | .136 | 1.580 | .116 |
| D.Bilirubin | -.289 | .159 | -.153 | -1.819 | .070 |
| Albumin | -.281 | .141 | -.146 | -1.998 | .047 |
| Glucose | -.594 | .141 | -.390 | -4.208 | .000 |
| AFP | -.072 | .108 | -.045 | -.670 | .504 |
| Hb | .156 | .151 | .096 | 1.035 | .302 |
| S.creatine | -.026 | .126 | -.014 | -.203 | .839 |
| BMI | -.039 | .116 | -.023 | -.335 | .738 |
| Alk.Phosph | -.022 | .164 | -.009 | -.135 | .893 |
| Gender | .065 | .103 | .041 | .635 | .526 |
Regarding VDR rs 7975232 gene polymorphisms, (A) allele was significantly higher in H. pylori infected group (75.6), while (C) allele was significantly higher in uninfected patients (85.6%). Regarding VDR rs 2228570 gene polymorphisms, (C) allele was significantly higher in H. pylori infected group (78.7%), while (T) allele was significantly higher in uninfected patients (74.1%).
This is inconsistent with the study of Martins et al., 2018 [20] who reported that the allelic and genotypic distribution analyses of the FokI and ApaI polymorphisms of the VDR gene did not show distribution differences between H. pylori-positive and negative groups.
The difference in the results may be attributed to the fact that infectious diseases, such as H. pylori infection, are the result of a complex interaction between genes and environmental and social factors. These interactions are crucial for the evolutionary processes and genetic composition of a population, which probably contributed (together with the ethnic influences) to the variation in the allelic frequencies observed in the study population.
Blood glucose was the only independent risk factor of rs 7975232. Meanwhile, blood glucose and s.albumin were the only independent risk factors of rs 2228570.
These results are similar to the study of Mackawy and Badawi, 2014 [21], who reported that FokI VDR was significantly associated with HOMA-IR, insulin, VitD, IL-6 levels in subgroup IIb. El Gendy et al., 2019 [22] also concluded associations which were found only between VDR FokI gene polymorphism and susceptibility to Egyptian patients with T2DM.
These results are in agreement with the study of Yassin et al., 2017 [23], who found that there was no significant association between (ApaI and TaqI) polymorphisms and anemia in terms of genotype and allele for the covariates (age, blood hemoglobin level, age at menopause (For women), educational level, ethnicity, medical history, serum ferritin and iron) but did not include blood sugar or s.albumin.
On the other hand, Mackawy and Badawi, 2014 [21] found that FokI VDR was significantly associated with Body Mass Index (BMI) in subgroup IIb, which is contrary to our results.
CONCLUSION
To our knowledge, this is the first study reporting a possible association between FokI and Apal VDR polymorphism and H. pylori infection.
RECOMMENDATION
Further studies on VDR gene polymorphisms and H.Pylori association, a potential study design, use of a large number of samples, and additional markers would improve the validity and reliability of findings.
LIMITATION OF THE STUDY
The sample size was limited. Therefore, large-scale studies are needed. Additional markers are urgently needed to be used to improve the validity and reliability of findings.
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
This study was approved by the ethical committee of the Tanta University, Egypt with approval number 31529/10/15
HUMAN AND ANIMAL RIGHTS
No animals were used in this research. All human research procedures followed were in accordance with the ethical standards of the committee responsible for human experimentation (institutional and national), and with the Helsinki Declaration of 1975, as revised in 2013.
CONSENT FOR PUBLICATION
Written informed consent was taken from each patient or relatives. Each patient received an explanation of the purpose of the study.
STANDARDS OF REPORTING
We confirm that we followed STROBE guidelines.
AVAILABILITY OF DATA AND MATERIALS
Not applicable.
FUNDING
None.
CONFLICT OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
Declared none.


