All published articles of this journal are available on ScienceDirect.
microRNA-146a: A Biomarker for Epileptogenesis, Epilepsy Prognosis, and Treatment Resistance
Abstract
Recently, more attention has been paid to identifying biomarkers for epilepsy to direct a more personalized treatment strategy, especially for patients who suffer from drug-resistant epilepsy which carries a much poorer prognosis. microRNA has emerged as an important and diverse type of biomarker that can participate in metabolic and cellular processes of the disease and, importantly, can be detected in patient’s serum. In this short review, we compile state-of-the-art evidence regarding miRNA-146a, a novel biomarker that shows high potential for studying epileptogenesis, monitoring disease progression, evaluating treatment response, and may even function as a therapeutic target given its role in the process of neuroinflammation.
1. INTRODUCTION
Epilepsy, which is characterized by recurrent unprovoked seizures, is one of the most severe neurological conditions and affects about 70 million people worldwide [1]. The disorder can be successfully treated in many cases with surgery or pharmaceutical intervention, however, more than 30% of patients will progress to drug-resistant epilepsy (DRE), which is generally synonymous with a poorer prognosis and quality of life, as refractory seizures will lead to significant neurological dysfunction, a need for more invasive treatment, and death in many cases [2, 3]. It is a decline in prognosis that presents a dire need for novel biomarkers that could alert clinicians of more complex diseases, and even predict it. Recently, more attention has been brought to microRNA alterations as a potential area of useful biomarkers for epilepsy. MicroRNAs are short, non-coding RNA molecules that are ubiquitous in the human body. They tend to contain complementary sequences to other coding genes, giving them the ability to silence, or inactivate an mRNA transcript they are a match for [4]. Importantly, microRNA molecules are present and detectable in serum, presenting an important noninvasive method of measuring potential biomarkers and therapeutic targets in a myriad of diseases.
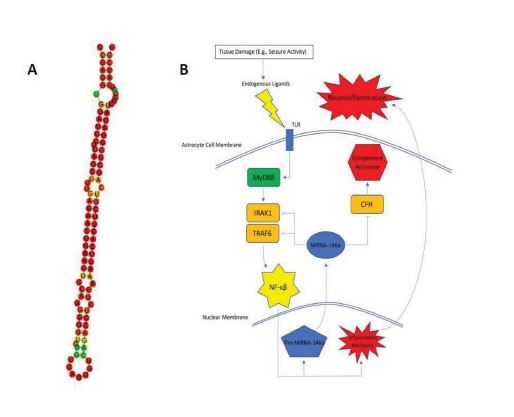
A promising candidate, miRNA-146a, has been documented as highly expressed by astrocytes and in areas of neuronal damage [5, 6]. miRNA-146a is a 99bp RNA molecule whose predicted secondary folding structure is visualized in Fig. (1A). In this short review, we will focus on recent literature characterizing miRNA-146a and its close relationship with epilepsy. We will assess its potential as a biomarker, discuss implicated pathways contributing to epileptogenesis and DRE, and address areas in need of further study.
The body of literature was unanimous that miRNA is upregulated in rat brains with induced temporal lobe epilepsy (TLE) compared to control groups [7-12]. This trend has since been replicated in human brain tissue [13-16]. The vast majority of studies focus on the hippocampus, which is a frequent site of epileptogenesis. Of all publications researching transcriptional changes of miRNA-146a, the only group that did not show a significant increase in miRNA-146a levels was Organista-Juarez et al. who assessed levels of miRNA in neocortical samples of brain tissue rather than hippocampal samples [17]. Although many of these publications assessed miRNA levels using brain tissue, studies have now shown that serum levels of miRNA, specifically miRNA-146a, closely mirror those found in brain tissue [10, 18, 19] and can be detected through a simple blood test. In particular, Ambrogini et al. [10] found that in rat models, serum increases of miRNA-146a mirrored the changes seen in hippocampal tissue. Additionally, An et al. [18] described significant increases in human serum miRNA-146a in a cohort of 90 patients with epilepsy when compared to the control group of non-epileptic individuals. This same study showed a positive correlation between serum miRNA-146a levels and NHS3 scores within the experimental group, suggesting a potential role of miRNA-146a as a prognostic biomarker in addition to a diagnostic biomarker alone.
Multiple studies have helped elucidate the category or categories of a biomarker that miRNA-146a falls into. Some researchers have studied aberrations in miRNA-146a levels in relation to the diagnosis and monitoring of epilepsy, such as seizure frequency, and seizure threshold [20]. Based on early studies on miRNA-146a in epilepsy, it is believed that it becomes upregulated in reactive astrocytes and in areas of neuronal damage in the acute to chronic stages of epilepsy development (epileptogenesis) [5, 6, 21]. Epileptogenesis is key in the development of the disease; having a biomarker that reliably informs clinicians that epileptogenesis is occurring would be monumental in the management and even prevention of epilepsy.
Considering the body of evidence, it now seems likely that levels of miRNA-146a could also be used to assess or predict epilepsies that are more resistant to pharmaceutical therapy. Two publications, in particular, showed a direct correlation between disease severity and levels of miRNA-146a. Organista-Juarez et al. showed a positive correlation between miRNA-146a levels and seizure frequency. Interestingly, there was also a positive correlation between miRNA-146a levels and the number of antiepileptic drugs (AEDs) that the patients were currently prescribed [17]. In another study, Leontariti et al. showed that there was significant upregulation of miRNA-146a in patients with known AED resistant epilepsies [14]. This paper also showed that there was a statistically significant clinical benefit to assessing the miRNA-146a level in these patients, further demonstrating its potential as a non-invasive, reliable predictor of epilepsy severity and potential for treatment resistance.
Current evidence strongly suggests that miRNA-146a plays a role in neuroinflammation [22-25]. In one of the first characterizations of miRNA-146a’s function, Taganov et al. found that miRNA-146a was endotoxin responsive [23]. Further analysis showed that miRNA-146a was induced by many microbial components and inflammatory cytokines. The team then utilized the Genomatix MatInspector software package to find complimentary binding sites that the miRNA could potentially target. They found consensus sequences on NF-KB and IRF3/IRF7. Using other prediction algorithms, they were able to find miRNA-146a target sequences within mRNAs encoding IRAK1, TRAF6, and COT/Tpl2/MAP3K8. IRAK1 and TRAF6 are intracellular proteins required for signal transduction from IL-1R1/TLR4 that leads to both transcription of inflammatory genes (secondary to NF-KB and AF-1 activation) and hyperexcitability of neurons (secondary to ceramide-induced calcium influx) [26].
Considering this established pathway and the gene silencing function of miRNA, it seems reasonable to conclude that miRNA-146a has an anti-inflammatory effect by acting as a negative control of the IRAK1/TRAF6 axis (Fig. 1B). Many published studies have echoed this conclusion [6-8, 27]. Wang et al. and Tao et al., in particular, showed evidence that the application of an miRNA-146a mimic can relieve seizures or delay seizure onset, respectively, in rat models. There is, however, some controversy regarding this conclusion. There have been multiple studies that provide evidence that silencing miRNA-146a can actually prove beneficial by alleviating neural damage [11, 28], protecting against status epilepticus [9, 29, 30], and even reversing drug resistance in DRE [12]. There were alternate pathways uncovered through these researches as well; for example: Zhang et al. found that when silencing miRNA-146a in a rat model of epilepsy, drug transport proteins associated with DRE (P-gp and MRP1) were found to be down-regulated [12]. Huang et al. found Notch-1 gene transcription to be significantly down-regulated after silencing of miRNA-146a [11] and Notch-1 signaling, leading to the excitation of CA1 pyramidal neurons during seizures [31]. Other anti-inflammatory pathways of miRNA-146a that have been suggested involve complement factor H (CFH) [22] and Bcl2/Bax [9].
| Author/Year | Methods | Results | Potential Biomarker Type |
| Huang/2022 | A case-control study. 80 patients with epilepsy and 70 healthy controls given MoCA, HAMA, and HAMD assessments followed by using qRT-PCR to assess levels of MiR-146a in the patients' serum. | micro-RNA-146a was significantly upregulated in patients with epilepsy when compared to control (0.808 (95%CI: 0.654 - 0.951, P < 0.001). Diagnostic potential using MiR-146a showed sensitivity of 87.8% and specificity of 68.2%. Levels of MiR-146a were negatively correlated with MoCa scores. | Diagnostic, prognostic |
| De Benedittis/2021 | A case-control study. qRT-PCR was used to measure the difference in human serum levels of MiR-146a between a cohort of 27 patients with TLE and a group of 20 healthy controls. | MiR-146a was significantly upregulated in TLE patient group compared to the healthy control group. Researchers found no correlation with disease severity. | Diagnostic |
| Leontariti/2020 | A case-control study. 162 patients with focal impaired awareness seizures. Clinical endpoint was DRE. qRT- PCR was used to measure serum levels of MiR-146a. | MiR-146a was significantly upregulated in patients with DRE compared to those without DRE, and suggested indepentant clinical value in predicting DRE (OR 3.076, 95% CI 1.467-6.452; P = .003). Authors also suggested that their analysis showed that evaluation of MiR-146a led to improved clinical benefit. |
Prognostic, treatment response |
| Martins-Ferreira/2020 | A case-control study. qRT-PCR was used to measure the difference in human serum levels of MiR-146a between a cohort of 79 patients with GGE and a group of 67 healthy controls. | MiR-146a was significantly upregulated in serum of the GGE group compared to healthy controls (3.13-fold). MiR-146a serum levels combined with MiR-155 and MiR-132 discriminated GGE from controls with an AUC of 0.85, 80% specificity, and 73% sensitivity. | Diagnostic |
| Organista-Juárez/2019 | A case-control study. qRT-PCR was performed on samples of brain tissue from two groups, 12 mTLE patients and 10 recently deceased controls to assess MiR-146a transcription levels. Further analysis compared transcription levels to markers of disease severity. | MiR-146a did not show significant differences in transcription between the mTLE and control groups. MiR-146a expression did show positive correlation with seizure frequency and number of AEDs prior to surgery. | Prognostic, treatment response |
| Elnady/2019 | Case-control study consisting of two groups: 30 children patients with epilepsy and 20 age-matched healthy controls. qRT-PCR was used to quantify expression levels of MiR-146a in serum of patients. Quantification of serum Ig was also performed. | Expression of MiR-146a was significantly upregulated in patients with epilepsy compared to control subjects (14.65-fold). There was no correlation found between MiR expression and Ig levels in patients' serum. | Diagnostic |
| Huang/2019 | A total of 128 rats were divided into a control group (N=20) and model group (N=108). The model group consisted of lithium-pilocarpine-induced TLE. Hippocampal tissue was subjected to qRT-PCR to assess expression of MiR-146a. This study also assessed the effect of modulating MiR-146a on epileptic events. |
MiR-146a expression was significantly higher in the TLE group when compared to the control group. Further, the team's results suggests that silencing MiR-146a led to alleviated neuronal damage in the hippocampus based on histological evaluation. |
Diagnostic |
| Ambrogini/2018 | Hippocampal tissue as well as serum of a small subset of 42 non epileptic rats and 48 induced-SE rats was subjected to qRT-PCR to assess expression of MiR- 146a. This study also assessed the effect of modulating MiR-146a on epileptic events. | MiR-146a expression was significantly upregulated in the hippocampi and serum of untreated SE rats when compared to the control group. The group also reported that administration of α-tocopherol negated differential expression of MiR-146a, suggesting that this MiR could be useful for assessing response to treatment. | Diagnostic, treatment response |
| Zhang/2018 | Hippocampal tissue of 40 rats with induced SE and 8 control rats were incrementally assessed for levels of MiR-146a using qRT-PCR at 1, 3, 7, 14, and 30 days. | MiR-146a expression was significantly upregulated in the SE group when compared to the control group at all time increments, with the most pronounced elevation seen at day 7 (approximately 5.5-fold change). This group's results suggest silencing MiR-146a led to alleviated neuronal damage in the hippocampus based on histological evaluation. | Diagnostic |
| An/2016 | Case-control study. Using qRT-PCR, the serum of 90 epilepsy patients and control populations were assessed for expression of MiR-146a using serum samples. | MiR-146a was significantly upregulated in epilepsy patients when compared to the control group. Further, MiR-146a levels were positively correlated with NHS3 scores, suggesting correlation with severity of seizures. Finally, the ROC of serum MiR-146a for prediction of epilepsy was 0.774, and increased to 0.887 when combined with serum MiR-106b levels. | Diagnostic, prognostic |
| He/2016 | Rats with induced SE and normal controls were used to study expression levels of MiR-146a in hippocampal tissue using qRT-PCR. SE-induced rats were sacrificed at 1 and 4-weeks post-SE induction. The group also used antagomir-146a injections to down-regulate MiR- 146a in a separate group. | MiR-146a expression was significantly elevated in SE-induced rats when compared to controls at both 1 and 4-weeks. Down-regulation of MiR-146a increased hippocampal expression of CFH and decreased seizure susceptibility, positing MiR-146a as a potential marker of epileptogenesis. | Prognostic, epileptogenesis |
| Matos/2014 | Male adult rats aged 60 days were randomly divided into a control group and a SE-induced group. Microarray analysis was used to reveal potential targets. qRT-PCR was then used to assess expression of MiR-146a in brain tissue. |
MiR-146a expression was significantly upregulated in SE-induced mice compared to the control group (FC: 1.99). | Diagnostic |
| Hu/2012 | Hippocampi of 102 rats (SE-induced and control) were used in statistical analysis for this study. Microarray analysis was used to assess potential MiR targets for further study. qRT-PCR was subsequently utilized to assess expression levels of MiR-146a. |
MiR-146a expression was significantly upregulated in the SE group when compared to the control group (Microarray: 2.69 FC, PCR: 2.5 FC). |
Diagnostic |
| Omran/2012 | Two groups of rats, 36 in the experimental group and 16 in the control group were used in this study. Epilepsy was induced in the experimental group using lithium-pilocarpine model. Additionally, brain samples of 5 children undergoing unilateral selective amygdalohippocampectomy for drug-resistant MTLE were also assessed. qRT-PCR was used to assess expression of MiR-146a. |
Significant upregulation of MiR-146a expression in hippocampal tissues of latant and chronic stages of MTLE in the rat model (higher in latent stage, mean of 2.8 compared to that of chronic stage mean of 1.8 FC). No significant difference was seen in the acute stage. In children with MTLE, MiR-146a was significantly upregulated when compared to control (FC of 3). |
Diagnostic, epileptogenesis |
| Iyer/2012 | 6 human brain tissue specimens from humans undergoing surgery with intractable epilepsy were contrasted with tissue from 6 age-matched controls with no history of seizures. qRT-PCR was used to assess expression of MiR-146a. | Significant upregulation of MiR-146a expression was seen in tissues of the experimental group when compared to the control group. | Diagnostic |
CONCLUSION
In conclusion, considering the BEST framework [32], we have evidence that miRNA-146a could be utilized as a diagnostic, monitoring, prognostic, or predictive biomarker (Table 1). There are many more miRNAs that have been described as aberrant in epilepsy; given that the scope of this paper is primarily to assess miRNA-146a’s usefulness as a biomarker, we did not conduct an exhaustive search and assessment of other miRNAs. Our assessment does, however, reveal that more work is needed to elucidate the complex role of miRNA-146a in the neuroinflammatory circuit. Whether its function is pro- or anti-inflammatory, though, there is great potential for miRNA-146a as a therapeutic target. miRNA has thus far proven a fascinating and fruitful area of biomarker research in many diseases, and we look forward to future research on the subject in relation to epilepsy.
LIST OF ABBREVIATIONS
| DRE | = Drug-Resistant Epilepsy |
| TLE | = Temporal Lobe Epilepsy |
| CFH | = Complement Factor H |
CONSENT FOR PUBLICATION
Not applicable.
FUNDING
None.
CONFLICT OF INTEREST
Dr. Liam Chen is the Editorial Advisory Board Member of The Open Biomarkers Journal.
ACKNOWLEDGEMENTS
Declared none.


